3CLpro Library
ChemDiv’s library of small molecules targeting 3CLpro contains 4,792 compounds.
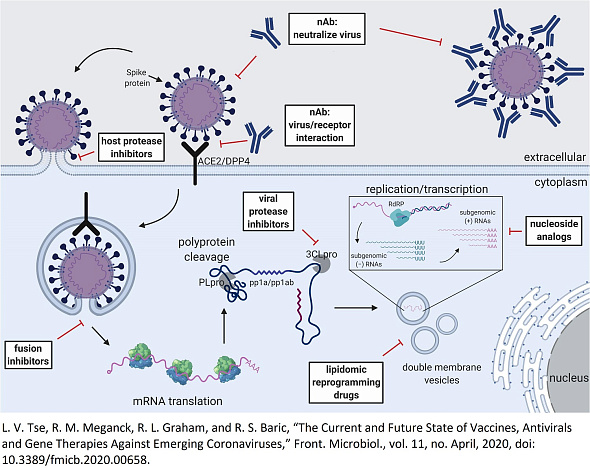
Coronaviruses are positive-sense, single-stranded RNA viruses characterized by large viral RNA genomes. Typically, beta-coronaviruses synthesize a polypeptide of approximately 800 kDa due to the transcription of their genome. This polypeptide undergoes proteolytic cleavage to produce various proteins essential for the virus's life cycle. The cleavage is mediated by two proteases: the papain-like protease (PLpro) and the 3C-like protease (3CLpro), also known as the 'main protease'. 3CLpro specifically cleaves the polyprotein at 11 distinct sites, generating various non-structural proteins crucial for viral replication. Unlike chymotrypsin-like proteases that use a serine residue in a catalytic triad, coronaviral 3CLpros utilize a cysteine residue in a catalytic dyad as the nucleophile. In solution, 3CLpro exists in an equilibrium between monomer and dimer forms. Each monomer comprises three structural domains: domains I and II, which include the catalytic site and a chymotrypsin-like scaffold, are connected to a third C-terminal domain via a long loop.
3CLpro is essential for the replication of coronavirus particles. It is located at the 3′ end of the viral genome, a region known for its high variability. The main protease itself exhibits conservation across different coronavirus species, making it an attractive target for therapeutics against existing and potentially emerging coronaviruses. Some natural compounds and their derivatives, known for anti-inflammatory and antiviral effects, have shown high binding affinity to 3CLpro, demonstrating its potential as a key element in antiviral drug development [1,2].
The development of 3CLPro inhibitors is a promising therapeutic strategy as this protease exhibits high conservation across coronaviruses, suggesting that these inhibitors could potentially be effective against a range of coronavirus strains, including future variants. Moreover, targeting a viral protease reduces the likelihood of impacting human cellular processes, minimizing potential side effects. Consequently, 3CLPro inhibitors are at the forefront of antiviral research, offering hope for effective treatments against current and emerging coronavirus-related diseases.
3CLPro targeted library is a subset of our SARS-CoV-2 small molecule collection.
The detailed methodology and related references for the SARS-CoV-2 library are comprehensively summarized in the corresponding slide deck. Specifically, the 3CLPro set was selected using a 3D shape similarity virtual screening approach, employing the Atom-Pair Fingerprint (APF) algorithm.
For the main portion of the 3CLPro subset, compounds were selected using the ligand for 4TWW PDB X-ray structure as the 3D APF template. The virtual screening process, which involves APF alignment, was conducted by setting the protein (i.e., the 4TWW PDB X-ray structure) as an excluded volume. This approach ensures that the selected compounds are complementary to the protein's binding site, avoiding regions bound to the protein. For a smaller portion of the 3CLPro subset, we utilized a series of designer molecules as 3D APF templates, developed in collaboration with Insilico Medicine. This strategy promoted exploration of novel chemical spaces, potentially leading to the identification of unique inhibitors targeting the 3CLPro enzyme of SARS-CoV-2
References:
[1] M. Tahir ul Qamar, S. M. Alqahtani, M. A. Alamri, and L. L. Chen, “Structural basis of SARS-CoV-2 3CLpro and anti-COVID-19 drug discovery from medicinal plants,” J. Pharm. Anal., Mar. 2020, doi: 10.1016/j.jpha.2020.03.009.
[2] D. Needle, G. T. Lountos, and D. S. Waugh, “Structures of the Middle East respiratory syndrome coronavirus 3C-like protease reveal insights into substrate specificity,” Acta Crystallogr. Sect. D Biol. Crystallogr., vol. 71, pp. 1102–1111, 2015, doi: 10.1107/S1399004715003521.