miRNA targeted small molecule library
Description
ChemDiv’s library of miRNA targeting molecules contains 20,000 chemically diverse compounds.
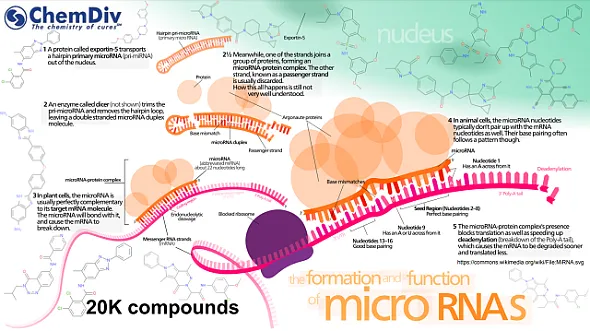
MicroRNAs (miRNAs or miRs) are small, approximately 18 to 25 nucleotides in length, non-coding RNAs that play crucial roles in the regulation of cell division, proliferation, and apoptosis. miRNAs are implicated in a broad spectrum of diseases, including cancer, neurological disorders, autoimmune diseases, metabolic syndromes, and cardiovascular diseases. Since their initial discovery in 1993, more than 2500 human mature miRNAs have been identified. The understanding of their mechanisms of action and functions remains a key focus of ongoing research [1,2].
Although numerous small molecules have been identified in the past decade, the exploration of miRNA-targeting small molecules is relatively nascent. We have created CheDiv's miRNA-targeted library, which we hope will contribute significantly to the advancement of this emerging field.
The library of small molecule compounds targeting miRNAs for drug discovery is an innovative and meticulously curated collection designed for exploring the therapeutic potential of modulating miRNA activity. This library stands out due to its specific focus on targeting miRNAs, which are key regulators of gene expression and are implicated in a variety of diseases
The molecule selection process was implemented using substructure Filters: The selection of molecules was rigorously conducted using substructure filters to exclude compounds identified as Pan-Assay Interference Compounds (PAINs) and those not adhering to SMARTS patterns. This filtering ensures the removal of reactive or otherwise undesirable molecules, enhancing the quality of the library.
To provide a high level of structural diversity, the library was constructed using a Hierarchical clustering approach coupled with the Min-Max algorithm, a method that ensures a wide structural diversity among the selected compounds. This diversity is crucial for exploring a broad range of potential interactions with miRNA targets.
ECFP-Tanimoto and Shape Similarity was implemented for further refinement, and similarity searches were conducted using ECFP combined with Tanimoto similarity measures alongside shape similarity assessments. These methods enable the identification of compounds with structural features and shapes likely to interact effectively with miRNA targets.
This collection represents a significant tool for drug discovery, particularly for conditions where miRNA dysregulation plays a crucial role. The careful selection and diversity of the compounds enhances the chances of identifying effective miRNA modulators. By targeting the intricate network of miRNA-mediated gene regulation, these small molecules have the potential to lead to breakthroughs in the treatment of various diseases, offering new avenues for therapeutic intervention.
Publications
[1] Alles J, Fehlmann T, Fischer U, Backes C, Galata V, Minet M, Hart M, Abu-Halima M, Grässer FA, Lenhof HP, Keller A, Meese E. An estimate of the total number of true human miRNAs. Nucleic Acids Res. 2019 Apr 23;47(7):3353-3364. doi: 10.1093/nar/gkz097
[2] Van Meter EN, Onyango JA, Teske KA. A review of currently identified small molecule modulators of microRNA function. Eur J Med Chem. 2020 Feb 15;188:112008. doi: 10.1016/j.ejmech.2019.112008