NEW KRAS-TARGETED LIBRARY
ChemDiv’s new KRAS-Targeted Library contains 16,000 compounds.
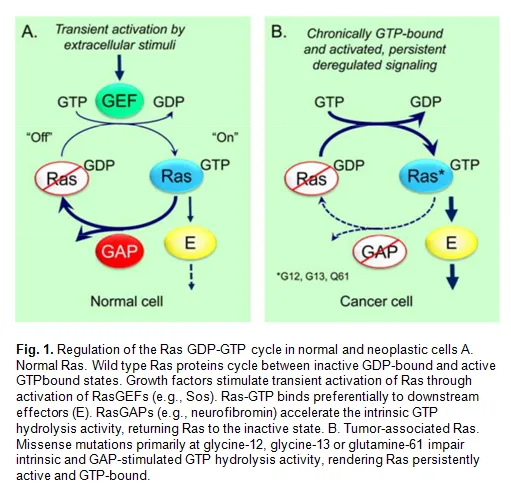
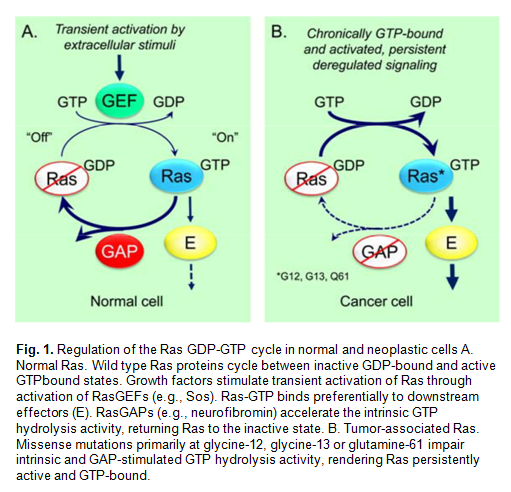
The small G-proteins KRas, HRas, and NRas function as GDP–GTP regulated binary switches in many signal transduction pathways that govern cell growth and differentiation. Alternation between the GDP-bound ‘off’ state and the GTP-bound ‘on’ state is controlled by the guanine nucleotide exchange factors (GEFs) SOS1 and SOS2, which catalyze the exchange of GDP for GTP (Fig. 1)
Gain of function mutations in Ras proteins are frequently found in human cancers and represent poor prognosis markers for patients. These mutations occur mostly in codons 12, 13, and 61 and render these forms resistant to GAP-catalyzed hydrolysis and chronically active.
Mutations of the KRas isoform constitute some of the most common aberrations among all human cancers and occur in three of the top four neoplasms that cause cancer deaths in the USA: lung, colon, and pancreatic cancer.
Binding sites and approaches
There are the following main approaches to inhibit KRas via binding to its surface:
- By blocking the Ras-effector interactions
- By trapping Ras in an inactive conformation incompatible with effector binding and signaling
KRAS G12C accounts for over 40% of all KRAS mutations, and has therefore been a key target for cancer drug developers. It is believed that covalent inhibitors may lock the KRAS G12C in the inactive state. These small molecules bind to a novel switch-II pocket. The following compounds have entered clinical trials in 2019:
- AMG 510 — Amgen
- MRTX849 — Mirati Therapeutics
- JNJ-74699157 (ARS-3248) — Johnson and Johnson
Recently small molecules bind to SI/II-pocket presented on both the active and inactive form of KRAS are described. These compounds inhibit the interactions between GTP-KRAS and the allosteric site of SOS1 as well as its effectors (CRAF and PI3Kα).
Researchers at Genentech, and other groups discovered small molecules that bind to a shallow pocket between the switch I and II regions of KRAS— switch I/II pocket (SI/II-pocket).
Virtual screening strategy
A combined profiling methodology that provides a consensus score and decision based on various advanced computational tools and theoretical knowledge:
- The common “MedChem” filters
- 2D-structure similarity to the reported K/HRas ligands (Tanimoto metric)
- Isosteric morphing procedure
- Privileged structures/fragments
- A set of compounds from PPI-targeted library (~5K compounds)
- A set of compounds with covalent warhead to target G12C mutant (~5K compounds)
More info ⬇️
https://store.chemdiv.com/catalog/focused_and_targeted_libraries/kras_targeted_library/
